The identification of a genetic susceptibility locus for systemic juvenile idiopathic arthritis (sJIA) has important implications for treatment of this severe, poorly understood illness, researchers reported.
Genetic analyses of a large, international cohort of sJIA patients determined that the IL1RN locus is associated with susceptibility to the disease and also influences response to treatment, according to Michael Ombrello, MD, head of the Translational Genetics and Genomics Unit, National Institute of Arthritis and Musculoskeletal and Skin Diseases, in Bethesda, Maryland, and colleagues.
High expression of IL1RN, which is genetically encoded to produce the acute phase protein IL-1RA, protects against the disease, while low expression of the gene and reduced production of IL-1RA confer susceptibility to the disease, they wrote in Arthritis & Rheumatology.
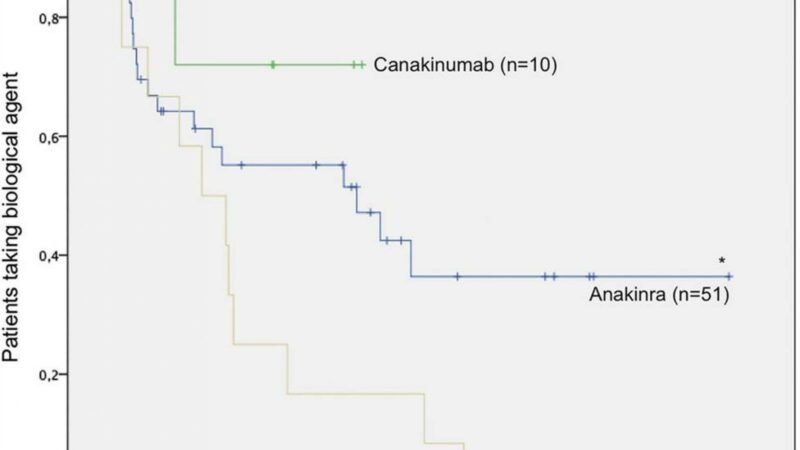
Moreover, patients who are homozygous for the high expression alleles of IL1RN SNPs are unlikely to respond to treatment with anakinra (Kineret), which is human recombinant IL-1RA.
sJIA is distinct from other forms of JIA, in that the chronic arthritis is accompanied by severe systemic inflammation, fever, rashes, lymphadenopathy, and hepatosplenomegaly. In many patients, that systemic phase ultimately resolves, but a damaging arthritis can persist for years. The current goal of treatment is to control the inflammation as quickly as possible during a potential window of opportunity that could prevent the arthritic sequelae.
The introduction of biologic therapies has profoundly improved the outlook in sJIA, but recent experience has shown that children vary considerably in their responses to biologics. Anakinra is typically used as a first-line treatment because of its rapid onset and other factors, but not all patients respond. Experience has shown that while some patients respond to anakinra and other agents that target interleukin (IL)-1, others respond to IL-6 blockers and still others to tumor necrosis factor inhibition. Being able to predict which children might respond to specific therapies would allow more rapid control over the systemic inflammation, the researchers noted.
In an accompanying editorial, Sebastiaan J. Vastert, MD, PhD, of University Medical Center in Utrecht, the Netherlands, and Peter A. Nigrovic, MD, of Harvard Medical School in Boston, summed up the significance of this study, stating, “A better grasp of disease pathophysiology, building upon the work of Ombrello and co-investigators, promises to open up a new era in pediatric rheumatology: the transition from one-size-fits-all treatment to tailored therapy. When replicated in a separate cohort, preferably in the context of prospective standardized treatment, the present study will be recognized as the first major step towards personalized medicine in sJIA.”
Gene Association Studies
In previous studies, researchers have utilized a candidate gene approach in an effort to identify genetic markers for sJIA, and reported that in small groups of patients they found more than 2 dozen SNPs at 11 susceptibility loci. In those studies, the researchers had looked at candidate genes that were of interest because of their function and potential importance in inflammatory pathways in general. But the genes that were identified were only weakly positive or ambiguous, and the results couldn’t be replicated in other populations. “In fact, all candidate gene studies across all diseases have been plagued by the same problem,” Ombrello told MedPage Today.
His group recently published the largest genetic study of sJIA, the International Childhood Arthritis Genetics Consortium case-control collection, which includes 770 children from nine countries. In that study, they identified two susceptibility loci and 24 additional possible sites, none of which overlapped with the previously identified sites in the candidate gene studies.
In the current study, they went back to the dataset for the 26 individual genetic markers that had been reported previously as being associated with sJIA and looked at the entire regions where those markers were located, extracting several hundred kilobases surrounding those loci. They therefore looked at 500 independent genetic markers, and found one that exceeded their relatively stringent significance threshold. That was the IL1RN region, and it contained seven strongly associated SNPs located in the promoter region of IL1RN that are inherited as a single haplotype. The SNP with the strongest link with IL1RA protein levels was rs4251961.
They further revealed that sJIA-associated SNPs correlated with expression of the IL1RN gene, and that variants that were protective against sJIA correlated with high expression of IL1RN, whereas those that were associated with risks for sJIA correlated with lower expression of IL1RN.
“Given that the association signal of the IL1RN locus was within the promoter region, we hypothesized that these SNPs may influence sJIA risk by altering gene expression,” they wrote.
The IL1RN locus also was interesting because of its potential relationship with treatment: Anakinra is the exogenous form of the endogenous protein encoded by IL1RN.
Response to Anakinra
“Given that the response of sJIA to treatment with recombinant human IL-1RA (anakinra) is variable, we hypothesized that individuals with the highest genetically encoded levels of IL-1RA may fail to respond to anakinra treatment more often than those with lower genetically encoded levels,” the researchers explained.
They assessed clinical responses and SNP genotype data from 38 children in the U.S. who had received anakinra, of whom 29 had shown any response, while nine were nonresponders.
For each of the seven IL1RN SNPs identified in the genetic analysis, patients who were homozygous for the high-expression alleles of IL1RN were associated with nonresponse to anakinra. This homozygosity for the high-expression alleles predicted a failure to respond with a sensitivity of 92% and a specificity of 71%.
Then, to assess whether the association of these SNPs and lack of treatment response was specific to anakinra or was a more general treatment recalcitrance, they did a similar analysis for 14 children who had been treated with tocilizumab (Actemra), which targets IL-6. They found no association with the previously identified SNPs and response to this agent. They also observed that eight of the 14 had previously been treated with anakinra without success, and that six did respond to tocilizumab.
“Taken together, these facts support the hypothesis that sJIA-associated IL1RN SNPs specifically predict nonresponse of sJIA to anakinra treatment, as opposed to identifying individuals whose sJIA is more broadly refractory to treatment,” the researchers wrote. “This is the first candidate biomarker that can prospectively guide therapeutic decision-making in sJIA,” they stated.
“It just seemed to make sense that while the people who are genetically encoded to make the highest level of IL1RA have protection against sJIA, that maybe they wouldn’t be amenable to treatment with the exogenous form of IL1RA, because they’re already making high levels of the protein,” Ombrello told MedPage Today.
Beyond Rheumatology
Interest in anakinra and other IL-1-directed therapies has increased for other more common types of diseases including sepsis and diabetes, and more recently, in the secondary prevention of heart attacks with the IL-1Β monoclonal antibody canakinumab (Ilaris), as was seen in the CANTOS trial. That study included more than 10,000 patients with previous myocardial infarction and found significant reductions in recurrent MIs or stroke. In an additional analysis, canakinumab also was shown to help prevent lung cancer metastases and to decrease lung cancer mortality.
This raises the question as to whether biomarkers identified in sJIA as predictive of response to anakinra might also extend to other agents and other conditions. “We don’t have data to make a statement about this either way, but we hypothesize that these genetic markers will co-predict responsiveness to canakinumab,” Ombrello said.
If that turns out to be true, the question becomes if the markers could identify people who might benefit more from secondary prevention of stroke, heart attack, and lung cancer metastases or alternatively, if they could predict patients who were unlikely to benefit.
“On one hand, identifying the right patients is a way of prioritizing them to get the treatment, and on the other hand, these are very expensive treatments, so being able to identify those who wouldn’t benefit could provide substantial cost savings to our healthcare system,” Ombrello said.
The study was supported by the Intramural Research Programs of the National Institute of Arthritis and Musculoskeletal and Skin Diseases and the National Human Genome Research Institute of the NIH, Arthritis Research UK, the German Federal Ministry of Education and Research, the Val A. Browning Charitable Foundation, the Marcus Foundation, the Manchester Academic Health Sciences Center, and the National Institute for Health Research Biomedical Research Unit Funding Scheme.
Ombrello and co-authors disclosed relevant relationships with Novartis, Novimmune, Abbott, AbbVie, Amgen, Baxalta Biosimilars, Biogenidec, Bristol-Myers Squibb, Astellas, Boehringer Ingelheim, Italfarmaco, Janssen, MedImmune, Novartis, Novo Nordisk, Pfizer, Sanofi, Roche, Servier, Takeda, UCB, GlaxoSmithKline, Aventis, Schwarz Biosciences, SOBI, Medac Pharma, and Chugai.
Related Content
-
May 5, 2015
A retrospective review of 77 SoJIA patients revealed that 50-70% achieved inactive disease or remission…
-
November 26, 2019
A disease you’ve never heard of is becoming increasingly common and carries a…
-
August 8, 2019
A single-center cohort analysis shows that lung disease (LD) is increasingly seen in…
-
January 17, 2018
Since 2000, the German Biologics register (BiKeR) has prospectively enrolled children with juvenile…
-
January 1, 1970
Systemic onset juvenile idiopathic arthritis (sJIA) is a dramatic onset inflammatory condition marked…
-
November 9, 2021